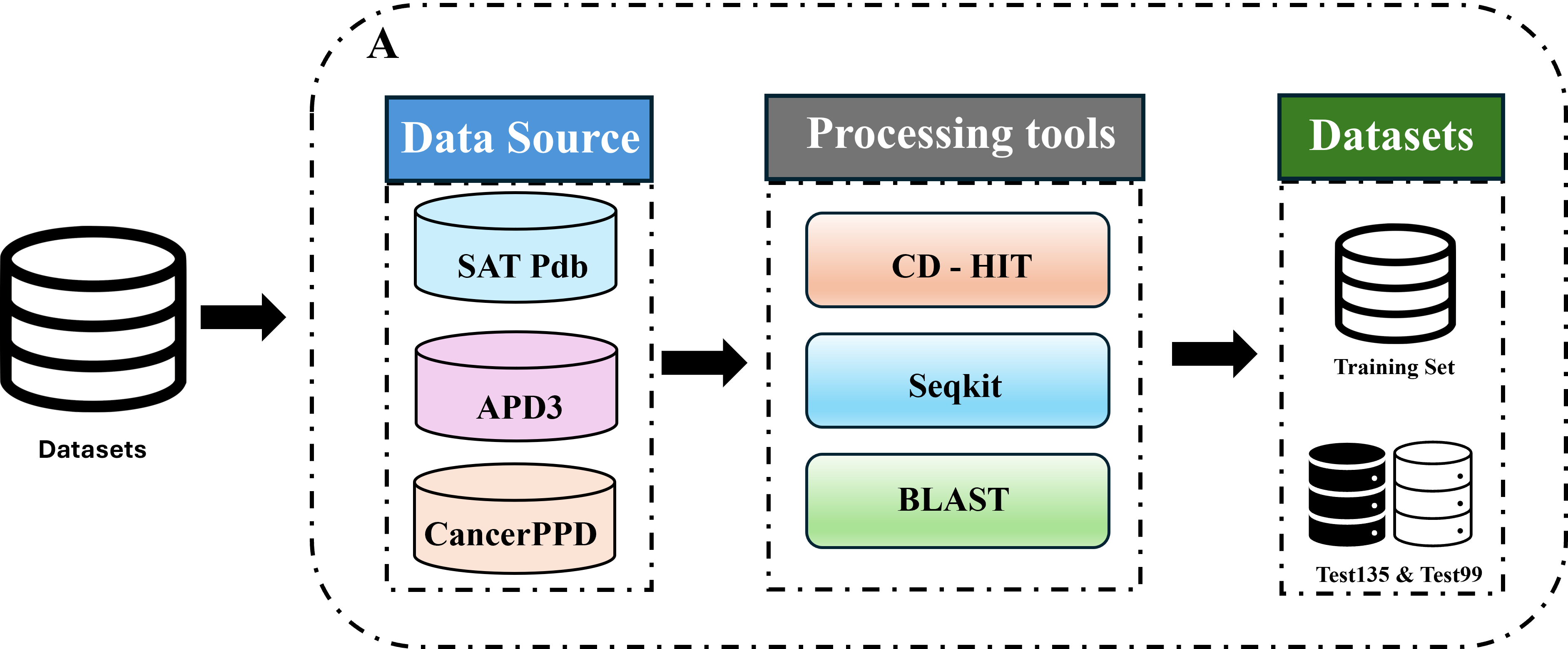
Part 1: Dataset.
The study initially collected sequences of anti-cancer peptides and non-anti-cancer peptides from multiple publicly available databases, such as SATPdb, APD3, and CancerPPD. Subsequently, a high-quality dataset was constructed through deduplication, length normalization, and cross-alignment utilizing tools like CD-HIT, SeqKit, and BLAST.
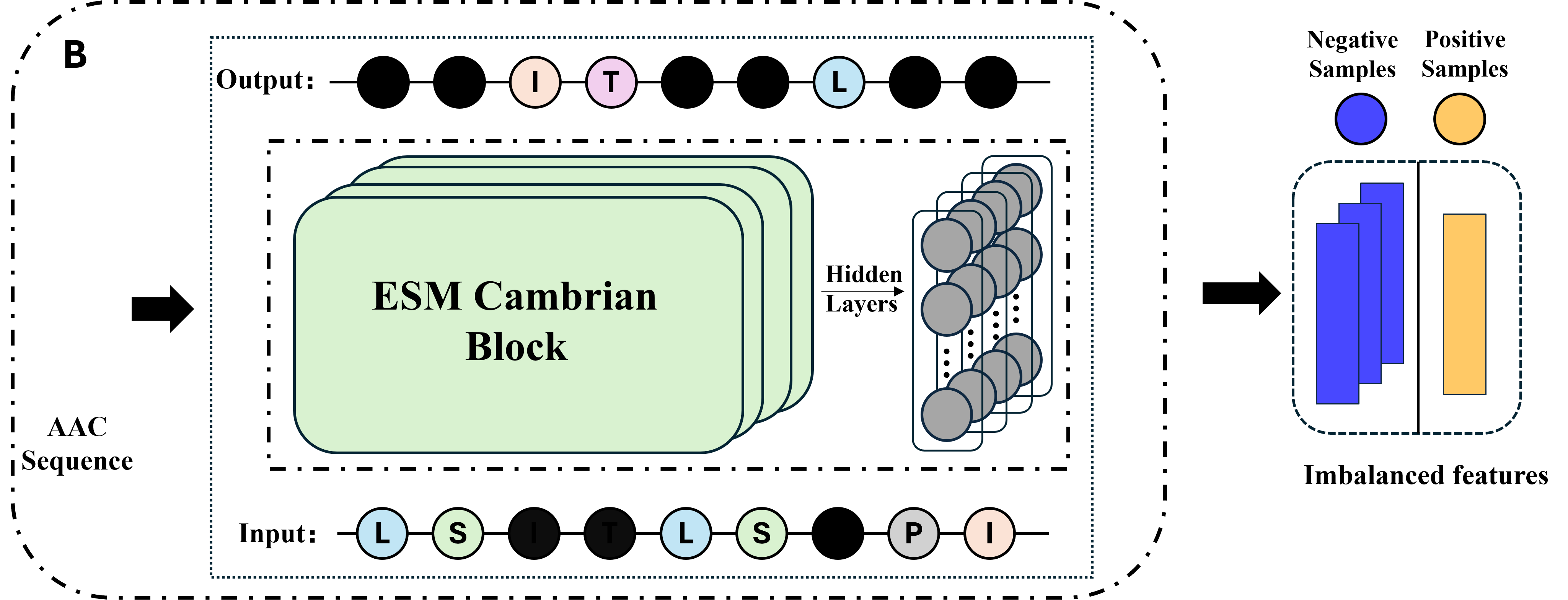
Part 2: ESMC representation learning module.
The ESMC pre-trained language model, which has been recently released, was utilized to conduct a deep featurization of peptide sequences. The ESMC model underwent unsupervised training on a corpus of hundreds of millions of protein sequences. This training allows it to effectively capture the contextual semantics, structural conformational information, and latent functional associations between amino acids. The high-dimensional vectors outputted by the ESMC model were employed to replace traditional statistical frequency and physicochemical property features. This substitution enabled a more comprehensive representation of sequence information.
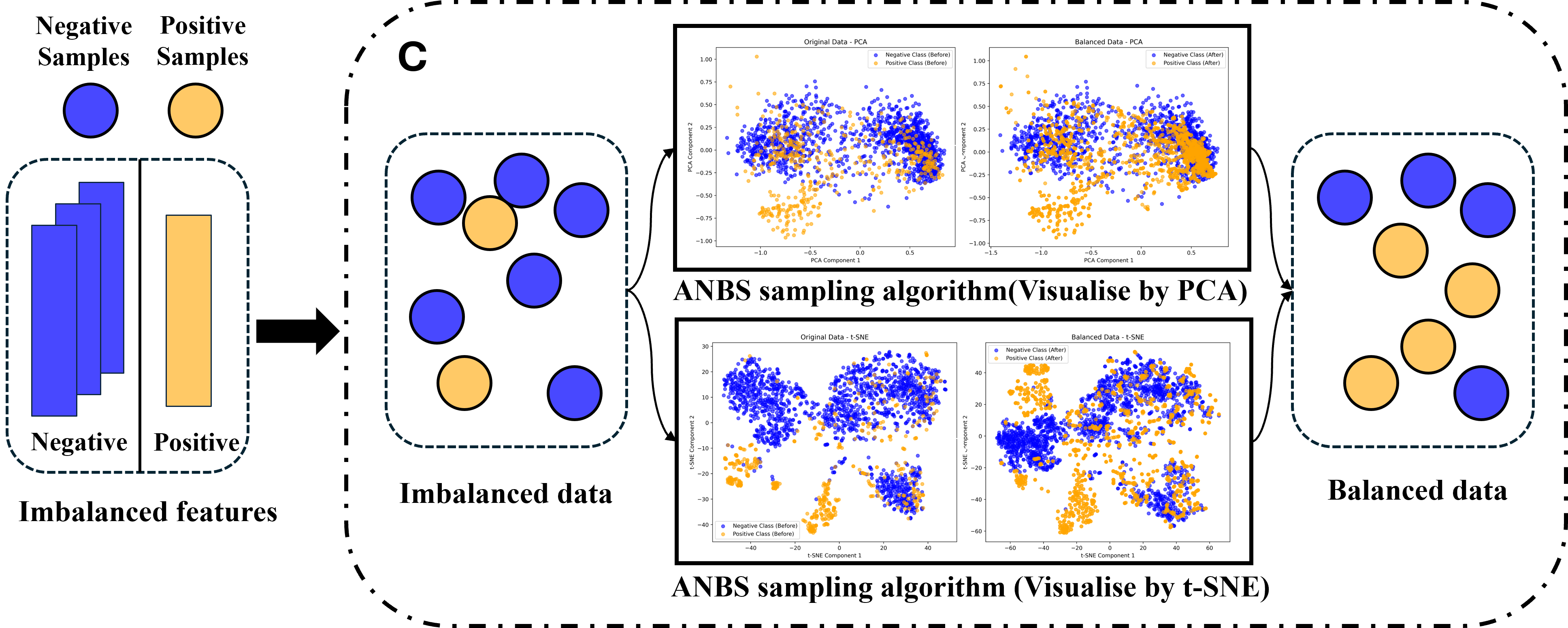
Part 3: Adaptive Neighborhood-Based Sampling (ANBS).
This paper presents a structure-aware oversampling method—Adaptive Neighborhood-Based Sampling (ANBS), the process of which is depicted in Figure 1C–D. By leveraging neighborhood structural information, this algorithm iteratively identifies majority-class samples near the decision boundary and subsequently synthesizes minority-class samples with discriminative value based on these identified samples. Consequently, ANBS enhances the model’s capability to recognize the boundary regions and circumvents the common issues of redundant augmentation and noise introduction that are typically encountered in traditional sampling methods.
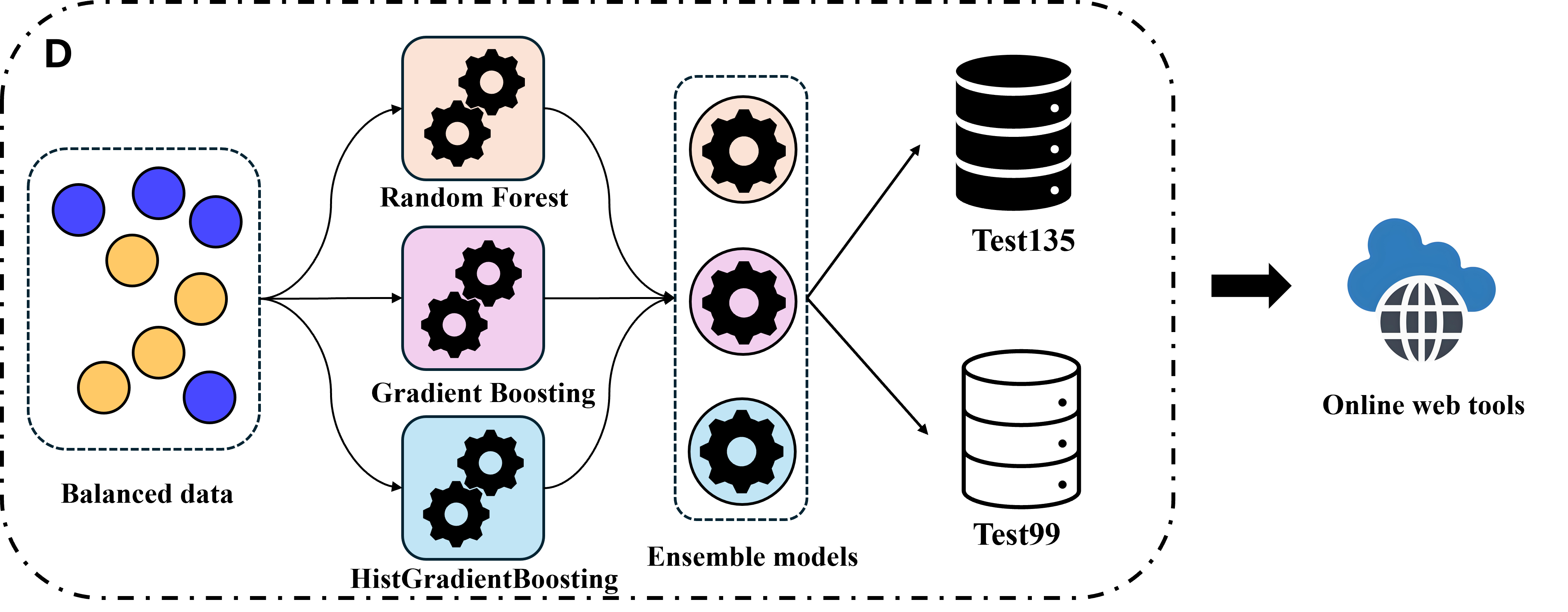
Part 4: Ensemble classifier.
The balanced data were subsequently fed into several base learners for training, and the strengths of different models were integrated through an ensemble strategy.
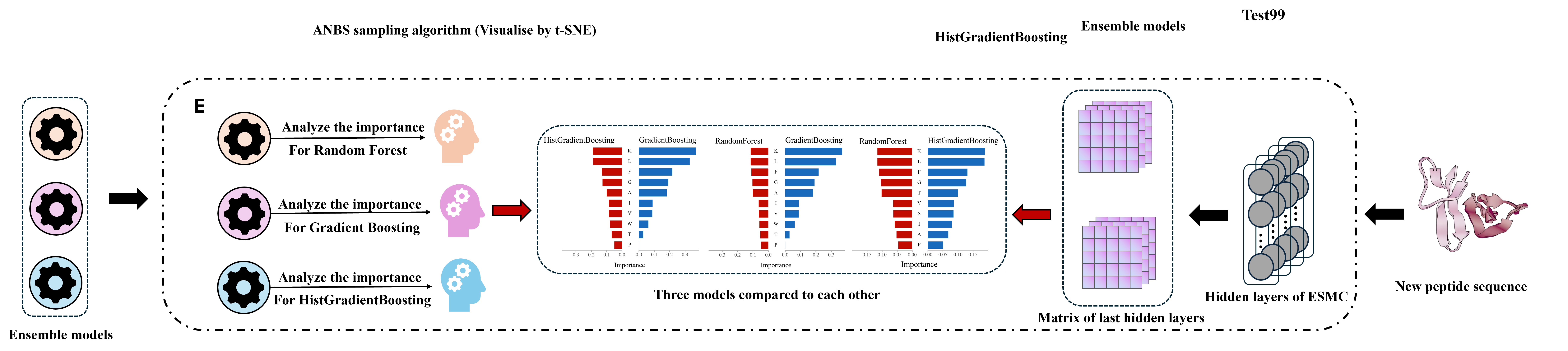
Part 5: Final evaluation and refinement module.
In this study, we further incorporate a feature importance analysis across multiple models to systematically compare the response mechanisms of Random Forest, Gradient Boosting, and HistGradientBoosting models. As demonstrated in the figure, the analysis results reveal that these models consistently focus on key residues such as K, L, F, and G. These residues are predominantly hydrophobic, basic, or flexible in their physicochemical properties. In addition, they have been confirmed to be closely related to anticancer activity in numerous studies.